Using gene expression fluctuations to understand and control biological systems

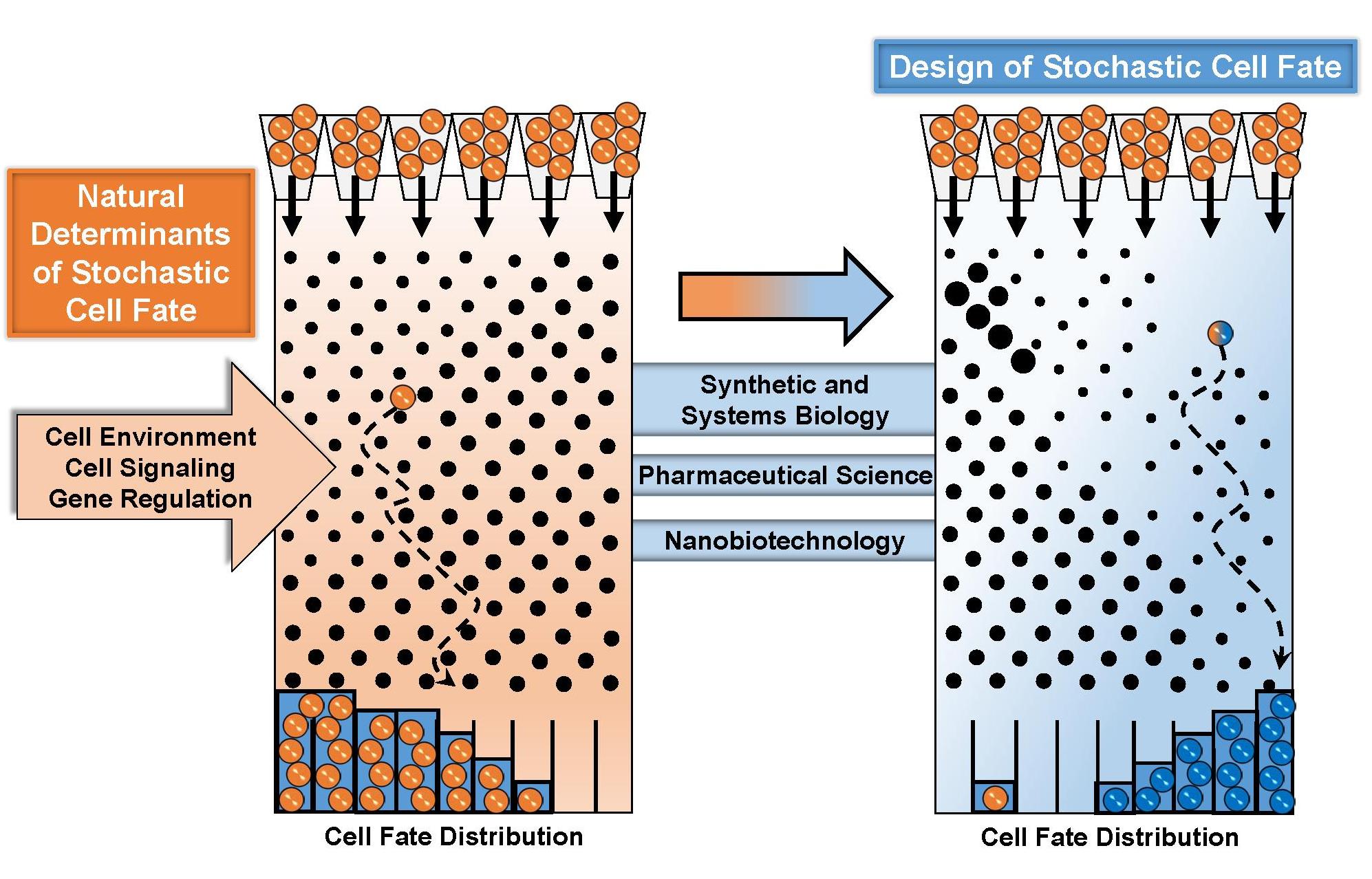
(Left) Monitoring single-cell decision making of latent HIV. [Adapted from Bohn-Wippert et al., Cell Reports (2018) (Above) Stochastic design of natural determinants of cell fate. [Adapted from Dar and Weiss, APL Bioengineering (2018)]
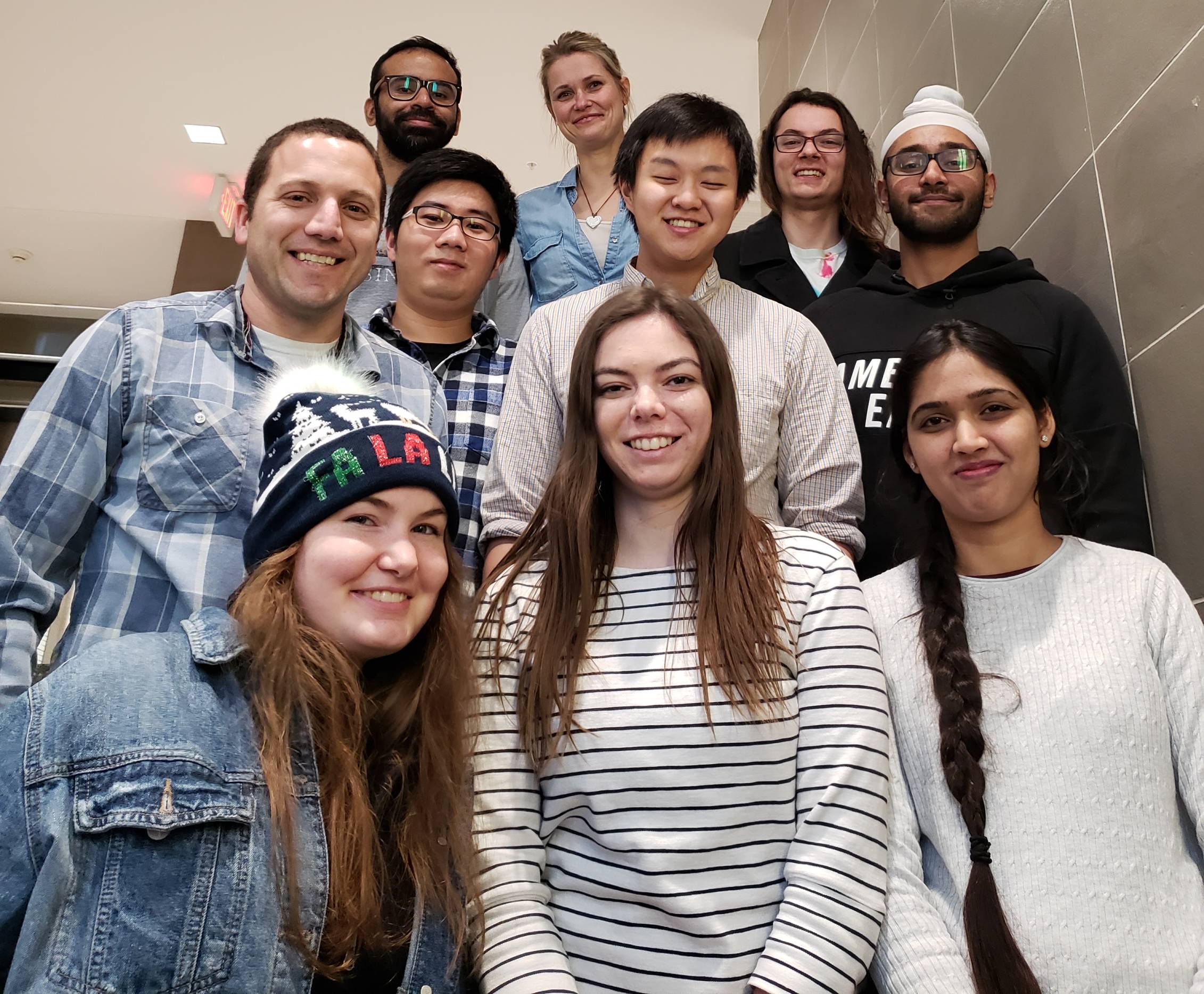
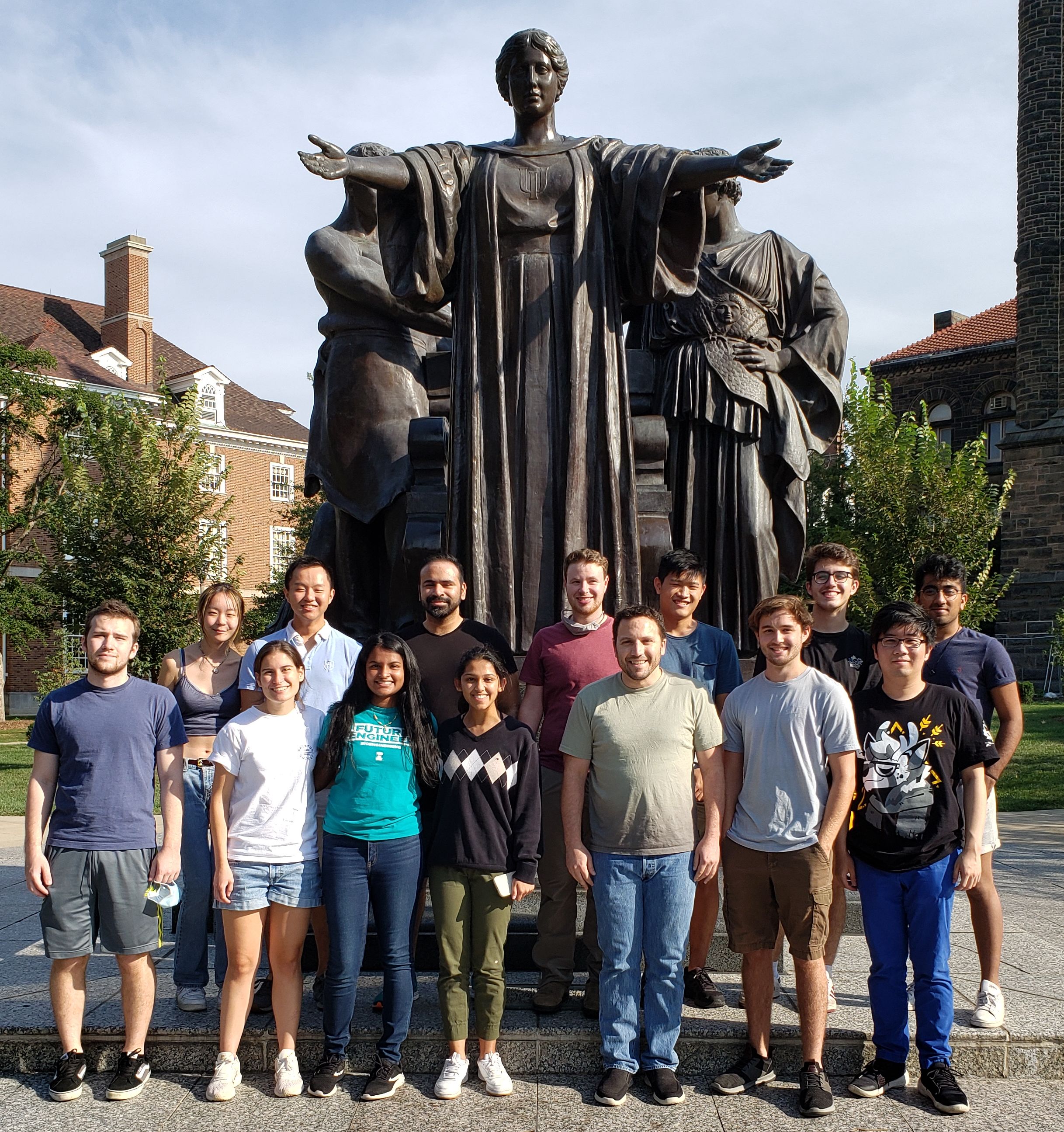
Group Photos December 2018 (left) and September 2021 (right)


